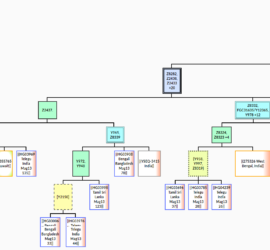
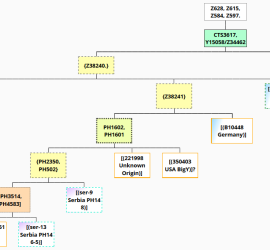
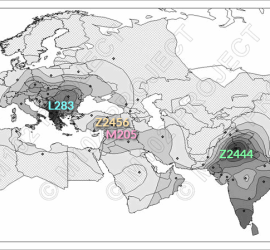
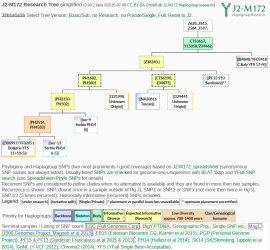
Y-SNP analysis of I4331, J2b2a-L283, Bronze Age Croatia – Mathieson et al. 2018
New DNA paper from Mathieson et al. 2018, The Genomic History of Southeastern Europe, gave us ancient DNA data from 225 individuals who lived in southeastern Europe and surroundings. Supplementary information from the paper states one of the sites analyzed was the Veliki Vanik burial mound, which is located near the town of Vrgorac in Split-Dalmatia County, in southern Croatia. […]