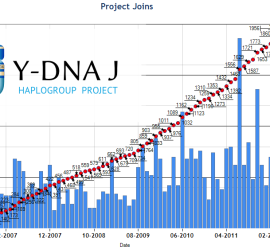
First Stats based on regrouping in J-Project@FTDNA (1,503 J2 kits). See Google J2-M172_spreadsheet > Stats Main Groups > 2014-01-06 (in order of importance): J2a1a1b1a1-M67 28.1%, J2a1a1a1-L24 23.5%, J2b2a1-L283 10.7%, J2a1b-Z6046 5.5%, J2a1a1a2-PF5197 5.4%, J2a1a1b-PF5116 (xM67,M319) 4,4%, J2a1a2-Z6055 4.1 %, J2a2-PF5008,L581 3.4%, J2a1a1b1a1-M319 2.7%, J2b1-M205 2.3%, etc.
Substantial new subgroups for Z467: S25258 and Z447,Z458(xL210)
As posted in the previous posts substream of the J2a-M67 haplogroup other new findings were possible lately. The most substantial refinement of the M67 tree was possible in the Z467 subclade using 8 BigY results, one of them (B1300/YF02101 Ireland) with additional YFull analysis, gre-77 Greek Hallast et al 2014, Sardinian #845 Francalacci et al 2015, 4 Genome of the […]